Object-oriented analysis
The CANlab tools enable interactive analysis of fMRI and other neuroimaging data through a small set of objects with simple, high-level methods (commands) tailored to neuroimaging. The toolboxes are distributed across linked GitHub repositories.
For the broader rationale behind interactive analysis, see the home page and the philosophy section.
Where to find walkthroughs and batch scripts
- Walkthroughs: the walkthroughs index is the main entry point. Source
.mfiles are in the CANlab_help_examples repository underexample_help_files; the rendered HTML reports are underpublished_html. - Batch scripts: the second-level analysis batch system lives in the same repository under
Second_level_analysis_template_scripts. See the batch system page for an overview.
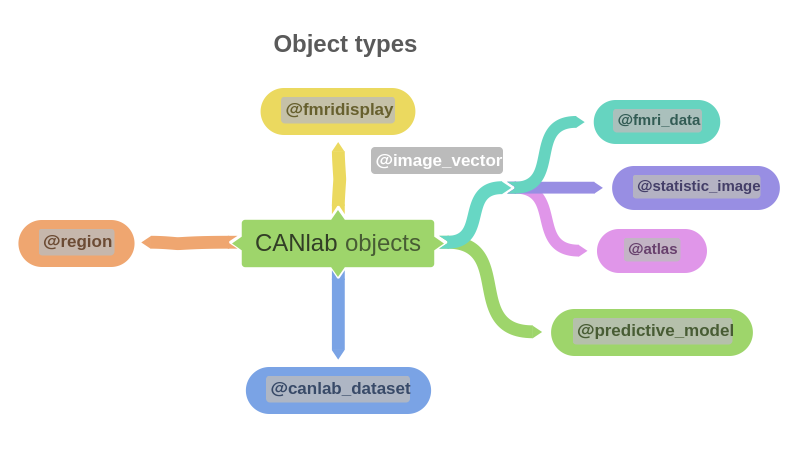
Types of objects
| Object | Description |
|---|---|
fmri_data | Stores neuroimaging datasets; you operate on a dataset by calling object methods |
statistic_image | Stores statistic images with stats and P-values |
atlas | Stores atlas data with probability maps and integers for unique regions |
region | Stores data grouped by region/network, treating each region as a unit of analysis |
fmridisplay | Container for handles to brain slices and surfaces, used for visualization |
fmri_data object: features and philosophy
- Flat format. Image data lives in a 2-D voxels × images matrix that’s space-efficient and easy to feed into any other analysis package.
- Compact storage. Out-of-image and empty voxels are removed, and data are kept in single precision.
- Reversible. Metadata is retained so you can reconstruct the 3-D image volume at any time. Built-in resampling makes it easy to combine datasets with different voxel sizes and bounding boxes.
- Multi-image. A single object can hold many images.
- Short, intuitive method names specialized for neuroimaging:
- Visualization —
plot,orthviews,surface,montage,histogram,isosurface - Image manipulation —
apply_mask,get_wh_image,resample_space,compare_space,flip,threshold - Data extraction —
apply_atlas,apply_parcellation,extract_gray_white_csf,extract_roi_averages - Analysis —
ica,mahal,image_math, and many more in thefmri_datasubclass
- Visualization —
- Provenance. History of changes to an object is tracked and updated.
- Documentation. In Matlab,
doc <class_name>(e.g.doc fmri_data) shows properties, methods, and examples.
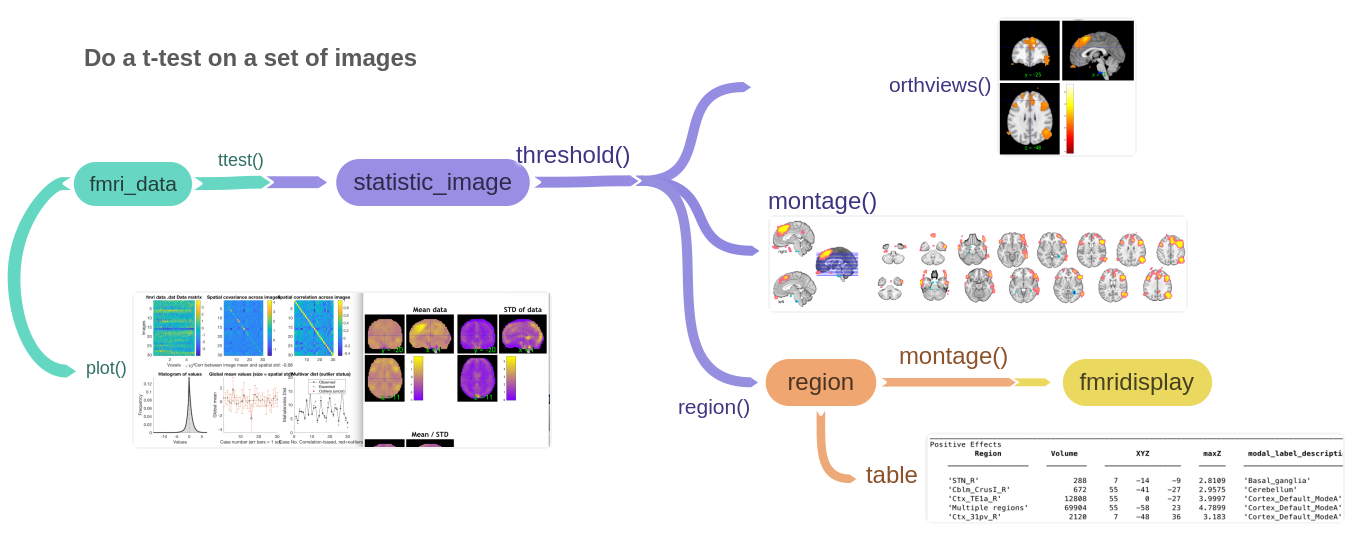
A simple analysis flow
The diagram below shows a flowchart for a simple group analysis: load a set of images (.nii or .img, one per subject) into an fmri_data object, then a few commands perform a t-test (or other analysis), threshold the resulting statistic map (a statistic_image object), and display interactive views, slice montages, region tables with automatic anatomical labels, and more.