Initial Setup and Requirements
This page walks through Matlab and GitHub setup and gives a fast on-ramp to the CANlab object-oriented tools.
If you just want to start, jump to the Quick-start walkthrough. The sections below explain the underlying requirements.
Requirements at a glance
| Component | What you need | Notes |
|---|---|---|
| Matlab | A recent release (R2019a or newer recommended) | Older releases may work but are not actively tested. |
| Matlab toolboxes | Statistics and Machine Learning, Signal Processing | Required by core CANlab functions. |
| SPM | SPM12 (free) | Used for image I/O and spm_orthviews. |
| Git | Any recent version | Optional but strongly recommended for updates. |
| Disk space | ~1–2 GB for the toolbox set, more for example datasets | Datasets in the walkthroughs may add several GB. |
Install Matlab
Install Matlab from the Mathworks website. Free trial versions are available. The CANlab tools require two add-on toolboxes:
- Statistics and Machine Learning Toolbox
- Signal Processing Toolbox
Some advanced walkthroughs (e.g. mixed-effects models) work better with the Optimization Toolbox as well, but it’s not required for the core walkthroughs.
Install Statistical Parametric Mapping (SPM)
The CANlab toolboxes use SPM’s image reading/writing functions and the spm_orthviews viewer. SPM is a free download. After unzipping, add the SPM folder to your Matlab path (addpath).
Install git and GitHub
Version control is optional here, but strongly recommended — it lets you pull updates with one command and contribute back. The fastest path:
- Install git.
- (Optional, friendlier UI) Install GitHub Desktop.
- New to git? Try the hello-world tutorial. For day-to-day flow see the GitHub flow guide and the git CLI essentials.
Install the CANlab core tools
Two options:
- Download a .zip file from each repository’s GitHub page (simplest, but you won’t get updates).
- Clone with git (recommended) — keeps you in sync with
git pull.
Start at github.com/canlab/CanlabCore:
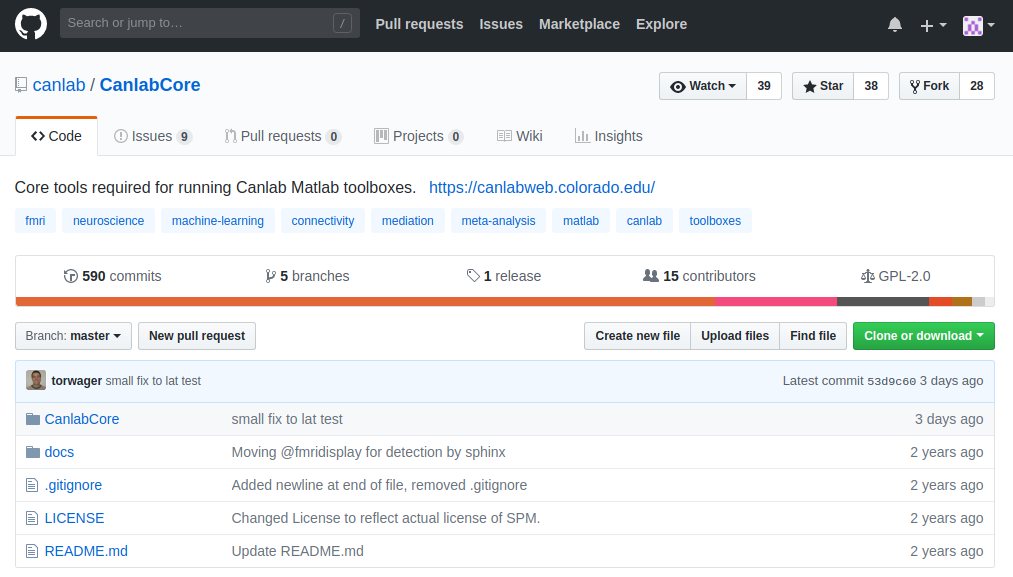
Use the green Code button to either download a zip or copy a clone URL. You can repeat this for the other repositories, or use the quick-start walkthrough below to grab everything at once.
Quick-start walkthrough
There is a Matlab script in the CanlabCore repository that downloads the repositories you need and adds them to your Matlab path. The walkthrough shows you how to use it:
Repositories
The CANlab imaging analysis tools are a set of linked repositories. Links and descriptions are on the repositories page.
Troubleshooting
| Symptom | First thing to check |
|---|---|
Undefined function or variable 'fmri_data' | CanlabCore not on path. Run which fmri_data — if empty, re-run the install walkthrough. |
spm_vol or spm_read_vols missing | SPM12 not on path. addpath(genpath('path/to/spm12')) and re-try. |
| Mac: “developer cannot be verified” on a SPM mex file | macOS quarantined the binary. From a terminal: xattr -dr com.apple.quarantine /path/to/spm12. |
| Path conflicts between SPM and other toolboxes | Run which -all <function> to see which copy Matlab finds first; reorder with addpath. |
| Walkthroughs fail to download example datasets | Check network/proxy, then try load_image_set from a fresh Matlab session. |
If you get stuck, please open an issue on the relevant repository’s GitHub page (most users will want CanlabCore issues). Include your Matlab version, OS, and the exact error text.